Many current diploid taxa used to be polyploid, but how can we tell?Whole-genome duplication is usually a cyclical process. Shortly after duplication, genomes begin to shed excess gene copies and restructure chromosomes, eventually returning to near pre-duplication genome sizes and chromosome numbers. This has made identifying ancient polyploids much more challenging than contemporary polyploids.
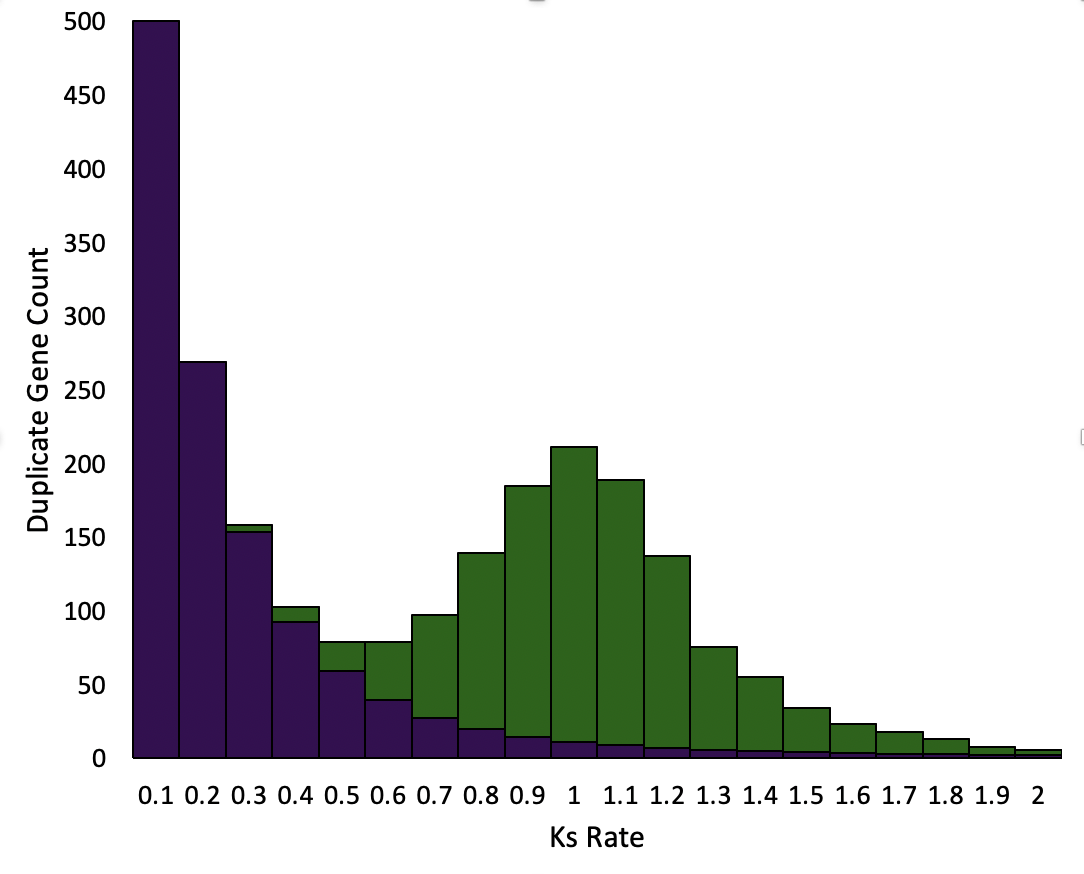
One technique used to assess ancient polyploidy involves identifying conserved gene duplicates. Even in highly-diploidized genomes, hundreds or thousands of genes are retained in duplicate--either due to selection or simply because sufficient time has not passed for them to be lost or differentiated. By measuring the synonymous substitution rate (Ks) between these retained duplicates, we can infer duplication events. All genomes have locally-duplicated genes with relatively short half-lives. When graphed together, these form a characteristic curve that falls quickly over evolutionary time (purple, at right). When the entire genome gets duplicated, a localized "pulse" of duplicate genes with similar Ks values is formed, and differentiates at roughly the same rate of time, created the classic secondary curve seen on Ks plots (green, at right). |
SLEDGE - Supervised Learning Estimation of Duplicated GEnomes
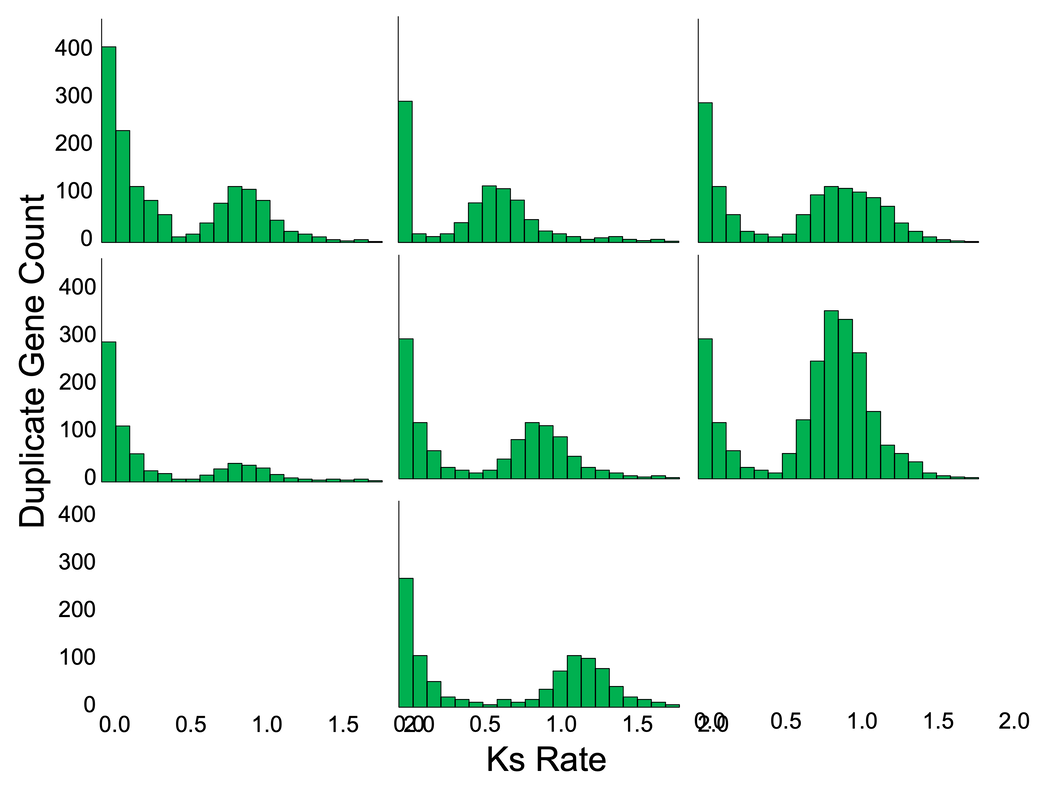
Working with collaborators George Tiley and Mike Barker, we designed a novel approach to assessing ancient genome duplication. Rather than explicitly modelling the evolution of specific genomes, we modeled the common features of Ks plots that either exhibit or lack evidence of whole-genome duplication. This approach allows for the rapid and computationally tractable screening of genome and transcriptome data for evidence of ancient duplications.
Current versions of SLEDGe have been tested against both simulated and real testing data, and perform well, correctly classifying 100% of genomes confirmed with syntenic data, 93.7% of all taxa in the 1000 Plant Transcriptomes dataset, and 87.4% of all taxa in the 1000 Insect Transcriptomes dataset. Preprint will be out soon! |